Research Articles
Graph Embedding for Mass Spectrometry Data Filtration: A Revolutionary Approach for Enhanced Metabolomic Analysis
This article explores the transformative potential of graph embedding techniques for filtering and analyzing complex mass spectrometry (MS) data, particularly in untargeted metabolomics.
Optimizing Microbial Cell Factories: Systems Metabolic Engineering for Next-Generation Biomanufacturing
This article provides a comprehensive overview of systems metabolic engineering strategies for developing high-performance microbial cell factories.
Navigating the Data Deluge: Key Challenges and Solutions in Untargeted Mass Spectrometry Metabolomics Data Mining
Untargeted mass spectrometry metabolomics generates vast, complex datasets, presenting significant bottlenecks in data mining that hinder biological insight and clinical translation.
Bridging the Gaps: Advanced Strategies for Completing Metabolic Networks in Biomedical Research
This article provides a comprehensive overview of gap-filling strategies for incomplete genome-scale metabolic models (GEMs), which are crucial for accurate metabolic simulation in biotechnology and drug development.
Overcoming Metabolic Heterogeneity in Cancer: Advanced Strategies for Research and Therapeutic Targeting
Metabolic heterogeneity is a fundamental characteristic of solid tumors that drives therapeutic resistance and complicates the development of effective cancer treatments.
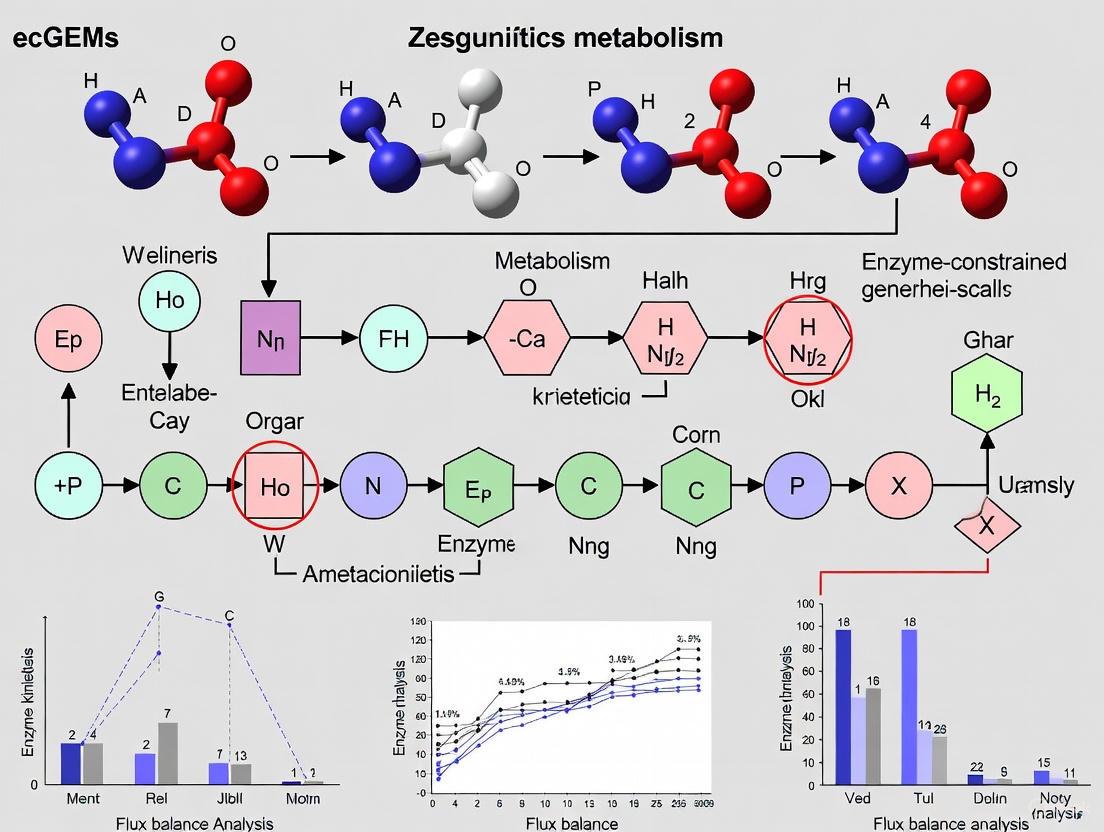
Enhancing Metabolic Model Accuracy: A Guide to Advanced kcat Prediction for ecGEMs
Enzyme-constrained genome-scale metabolic models (ecGEMs) have emerged as powerful tools for predicting cellular phenotypes, optimizing metabolic engineering, and understanding proteome allocation.
BMR Measurement Agreement: Evaluating Indirect Calorimetry vs. Predictive Equations in Clinical and Research Settings
This article provides a comprehensive analysis for researchers and drug development professionals on the agreement between Resting Metabolic Rate (RMR) measurements obtained via indirect calorimetry, the gold standard, and those...
Overcoming the Hurdles: How Machine Learning is Tackling Metabolic Pathway Optimization Challenges
This article reviews the significant challenges in optimizing metabolic pathways for microbial cell factories and drug development, and how machine learning (ML) is providing innovative solutions.
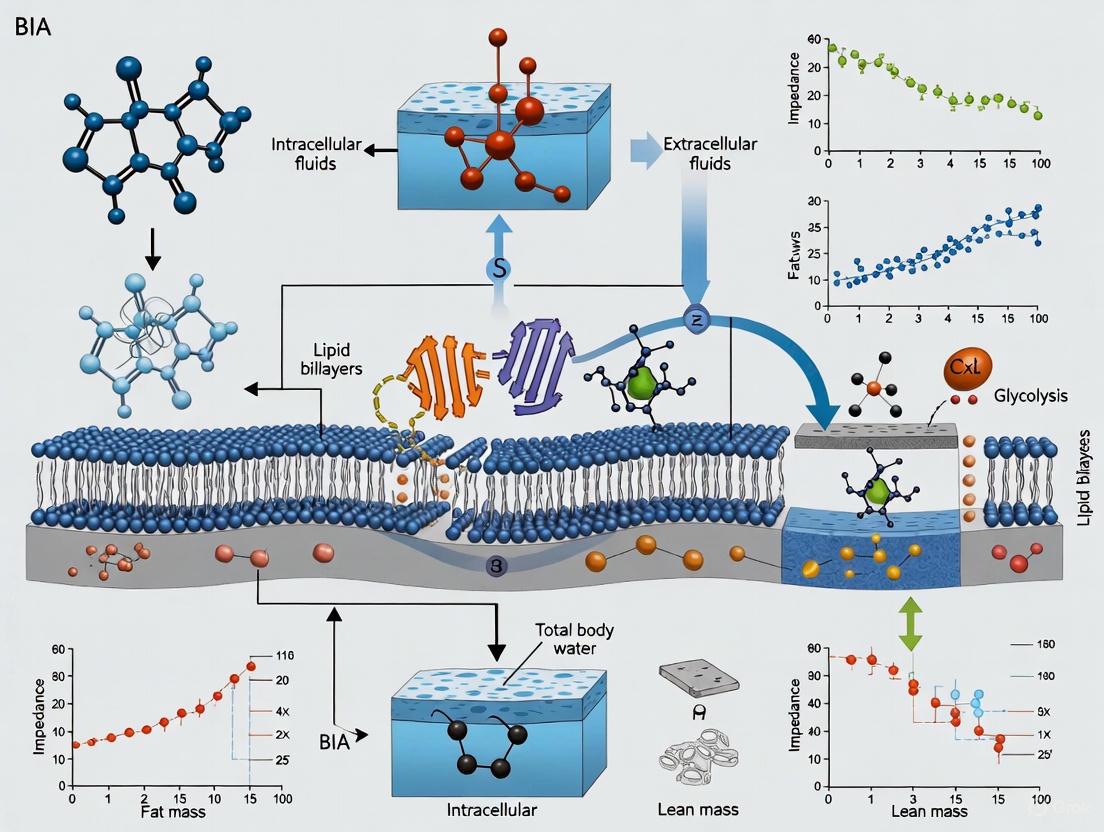
Bioelectrical Impedance Analysis (BIA) in Biomedical Research: From Body Composition to Advanced Drug Screening Platforms
This article provides a comprehensive analysis of Bioelectrical Impedance Analysis (BIA) for researchers, scientists, and drug development professionals.
Enzyme-Constrained Genome-Scale Metabolic Models: A Guide to Enhanced Predictions in Biomedical Research
Enzyme-constrained genome-scale metabolic models (ecGEMs) represent a significant advancement over traditional stoichiometric models by integrating enzyme kinetics and proteomics data to enhance the prediction of cellular phenotypes.